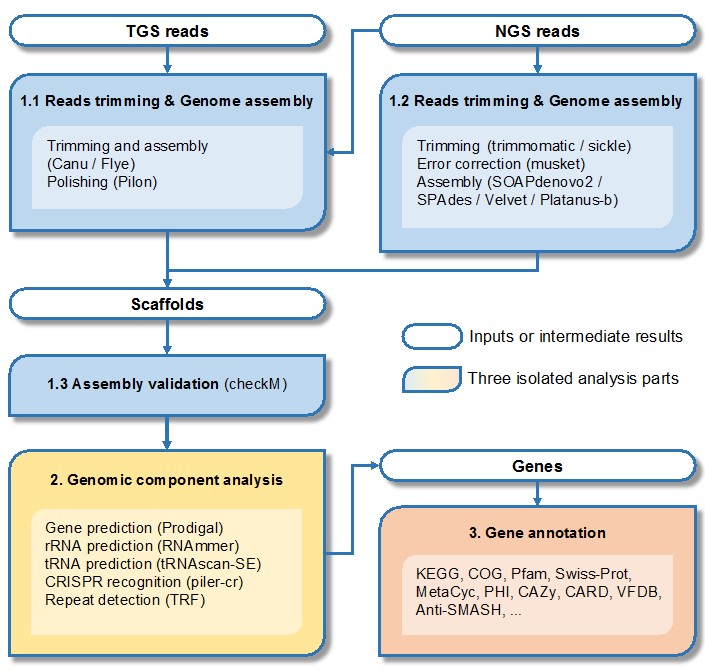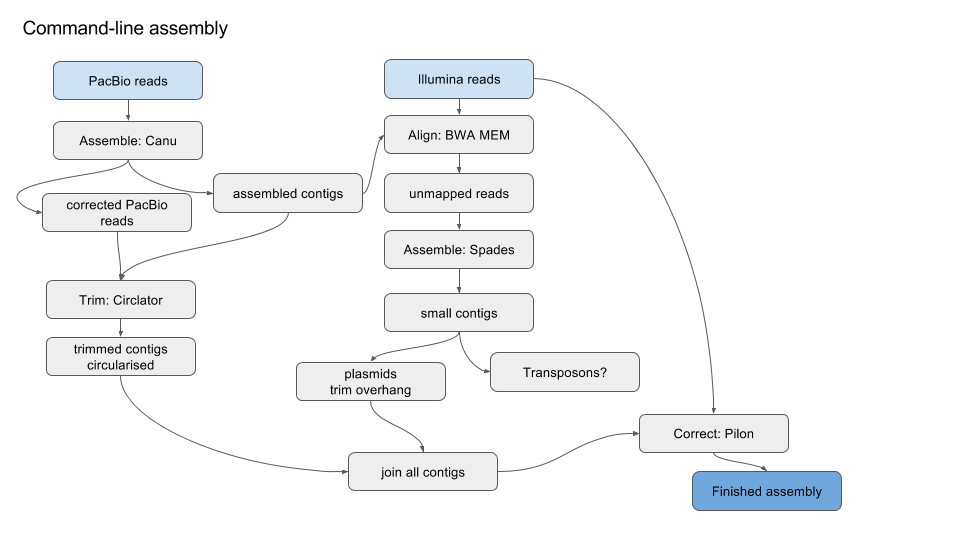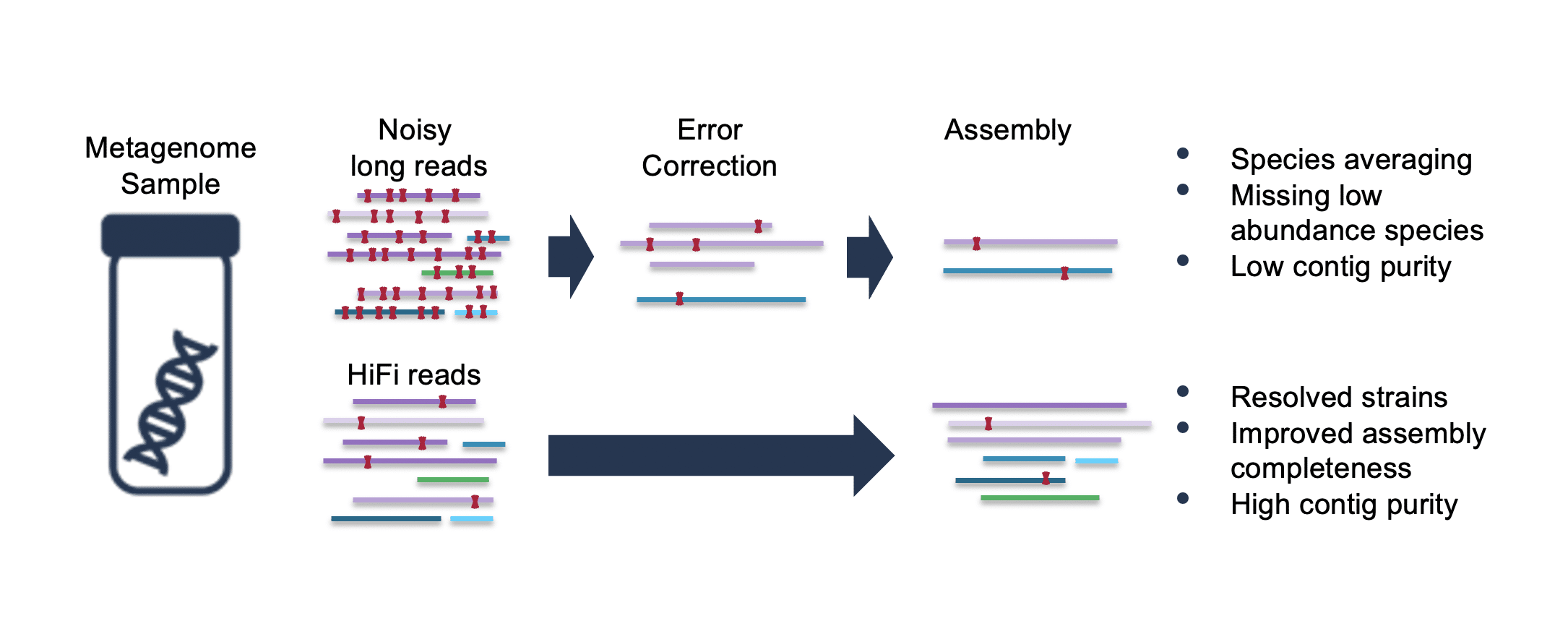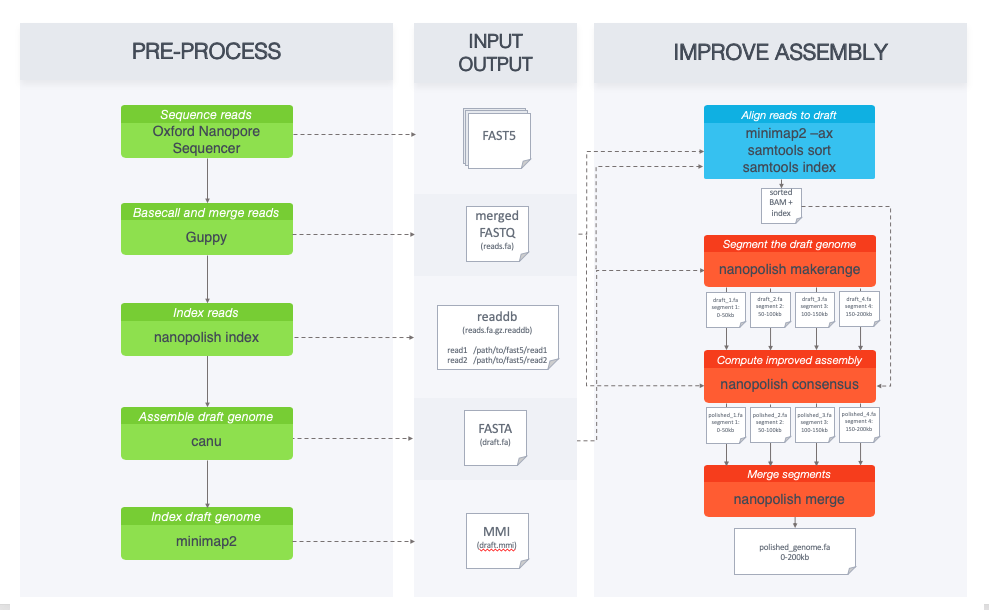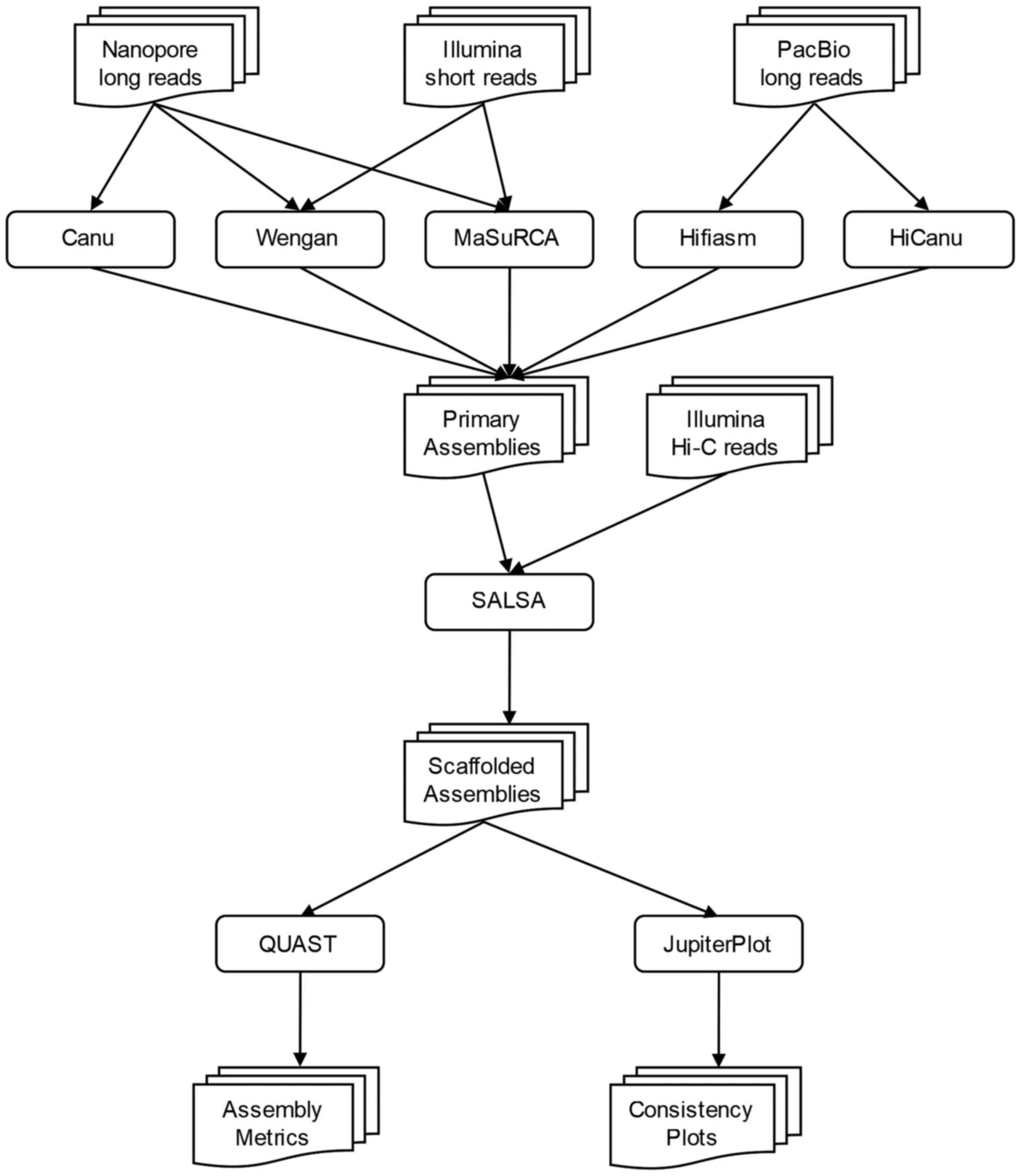
Benchmarking of next and third generation sequencing technologies and their associated algorithms for <em>de novo</em> genome assembly

A practical comparison of the next-generation sequencing platform and assemblers using yeast genome | Life Science Alliance
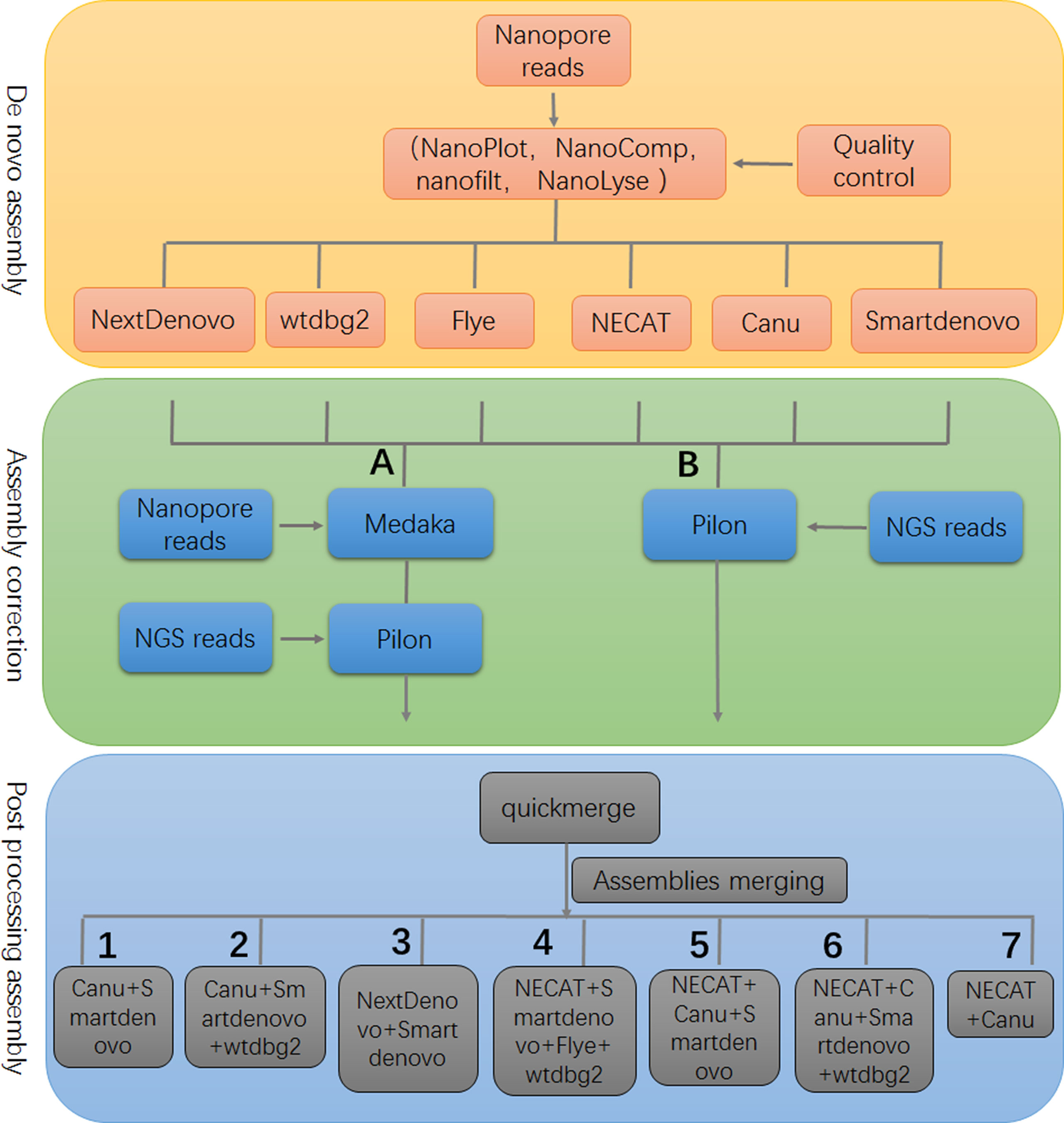
Frontiers | Systematic Comparison of the Performances of De Novo Genome Assemblers for Oxford Nanopore Technology Reads From Piroplasm
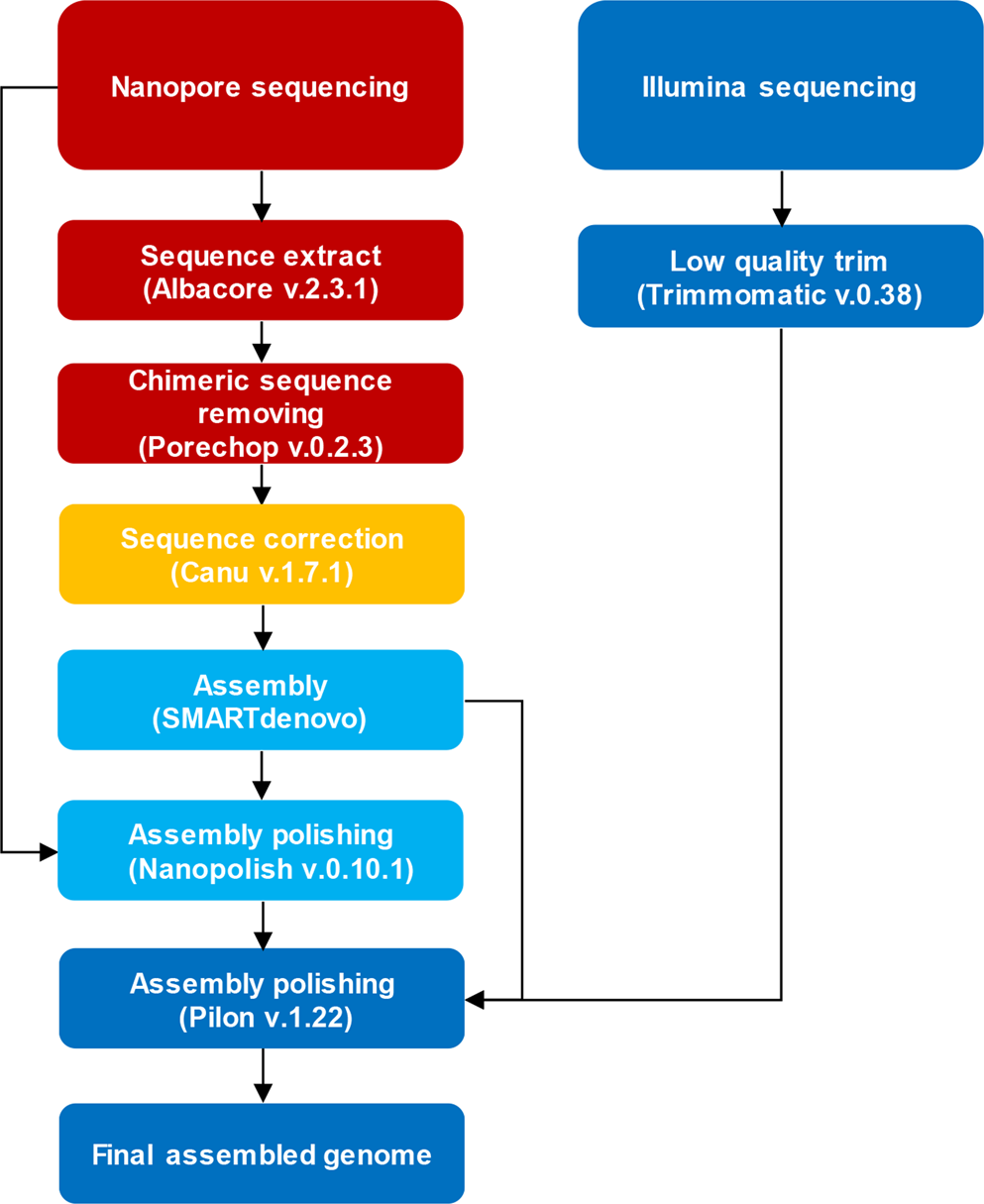
Nanopore sequencing reads improve assembly and gene annotation of the Parochlus steinenii genome | Scientific Reports

Comparative Evaluation of Genome Assemblers from Long-Read Sequencing for Plants and Crops | Journal of Agricultural and Food Chemistry

Comparisons between Canu and Flye assemblies. A, B. Dot-plots of the... | Download Scientific Diagram
Benchmarking of next and third generation sequencing technologies and their associated algorithms for de novo genome assembly

Highly accurate-single chromosomal complete genomes using IonTorrent and MinION sequencing of clinical pathogens - ScienceDirect
Frontiers | Comparing Long-Read Assemblers to Explore the Potential of a Sustainable Low-Cost, Low-Infrastructure Approach to Sequence Antimicrobial Resistant Bacteria With Oxford Nanopore Sequencing

The effect of coverage on Canu genome assembly contiguity. The number... | Download Scientific Diagram

Molecular Ecology on Twitter: "Attainable chromosome level genome assemblies for your favorite ecologically interesting model. A step-by-step approach so you too can generate a genome assembly for your organism(s)!!! @LucianoMatzkin #cactusfly Link

Raw pacific biosciences and illumina sequencing reads and assembled genome data for the cattle ticks Rhipicephalus microplus and Rhipicephalus annulatus - Data in Brief

An exploration of assembly strategies and quality metrics on the accuracy of the Knightia excelsa (rewarewa) genome | bioRxiv
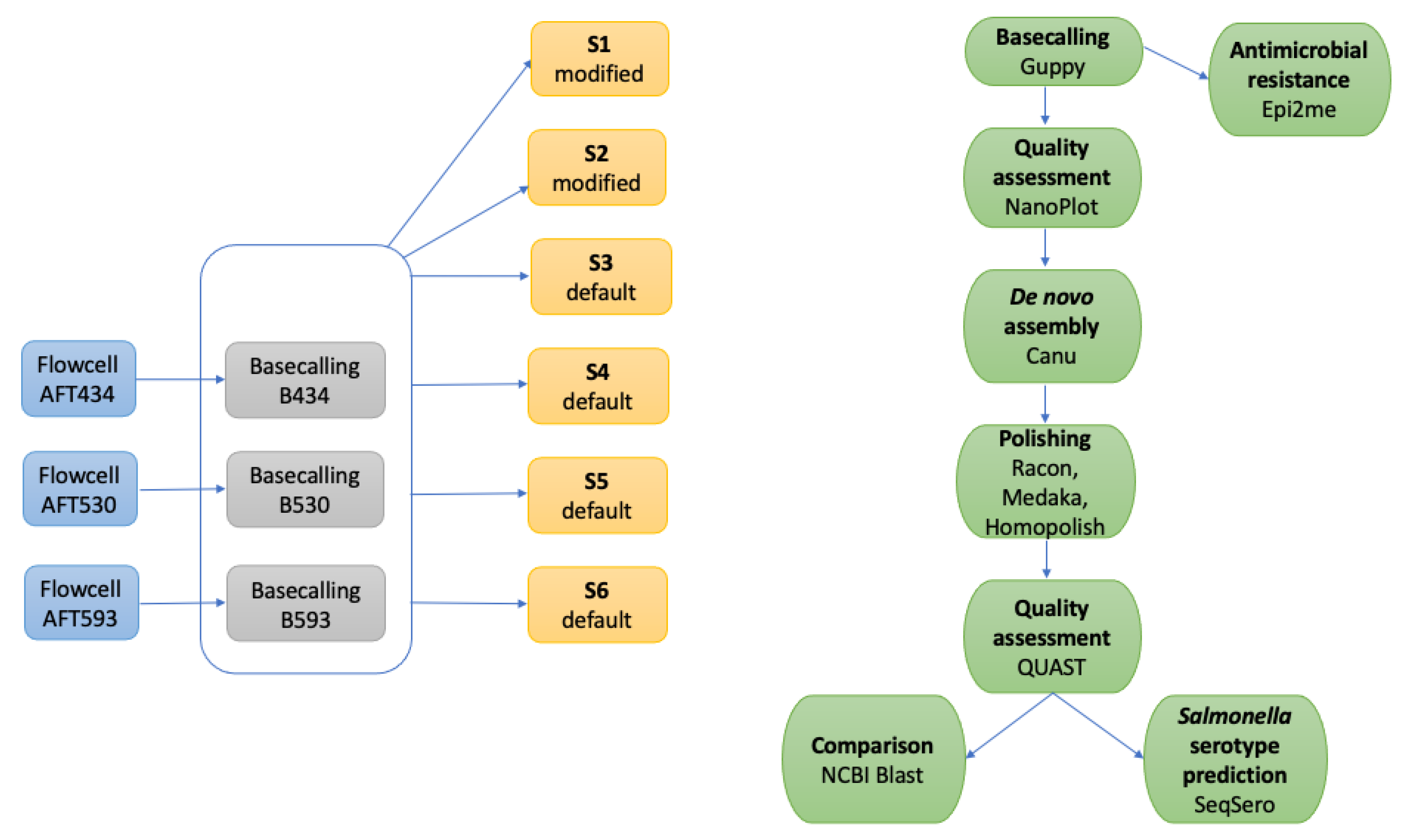
Applied Sciences | Free Full-Text | Factors Affecting the Quality of Bacterial Genomes Assemblies by Canu after Nanopore Sequencing